
Ajay Khanna
Bridging quantum mechanics and real-world impact through drug discovery, energy transfer, and sustainable technologies
Research Impact Areas
Computational chemistry at the intersection of molecular science and transformative applications
Drug Discovery
Virtual Screening & Design
Key Capabilities:
- Virtual screening pipelines
- Protein-ligand binding
- ADMET prediction (ML)
- Binding affinity calculations
Energy Transfer
FRET & Excitonic Coupling
Key Capabilities:
- FRET mechanisms
- Excitonic coupling calculations
- Light-harvesting systems
- Charge transfer dynamics
Energy Storage
Battery Materials
Key Capabilities:
- Electrode materials design
- Electrolyte optimization
- Li-ion migration pathways
- Solid-state batteries
Green Energy
Sustainable Solutions
Key Capabilities:
- Organic photovoltaics
- Catalytic CO₂ reduction
- Water splitting mechanisms
- Sustainable materials
About Me

Ajay Khanna, Ph.D.
Postdoctoral Researcher
Los Alamos National Laboratory
Research Objective
I'm a computational chemist at Los Alamos National Laboratory, specializing in the development and application of quantum mechanical methods and machine learning to solve critical challenges at the intersection of molecular science and real-world applications.
My research bridges fundamental theoretical chemistry with transformative solutions in drug discovery, energy materials, and sustainable technologies. I believe the most impactful computational research emerges at the intersection of rigorous quantum mechanics, efficient algorithms, and practical applications.
Current Research at LANL
Developing deep learning frameworks for non-adiabatic molecular dynamics simulations
Investigating chiroptical spectroscopy in π-stacked molecular aggregates
Designing QSPR models for energy transfer materials
Core Expertise
Projects
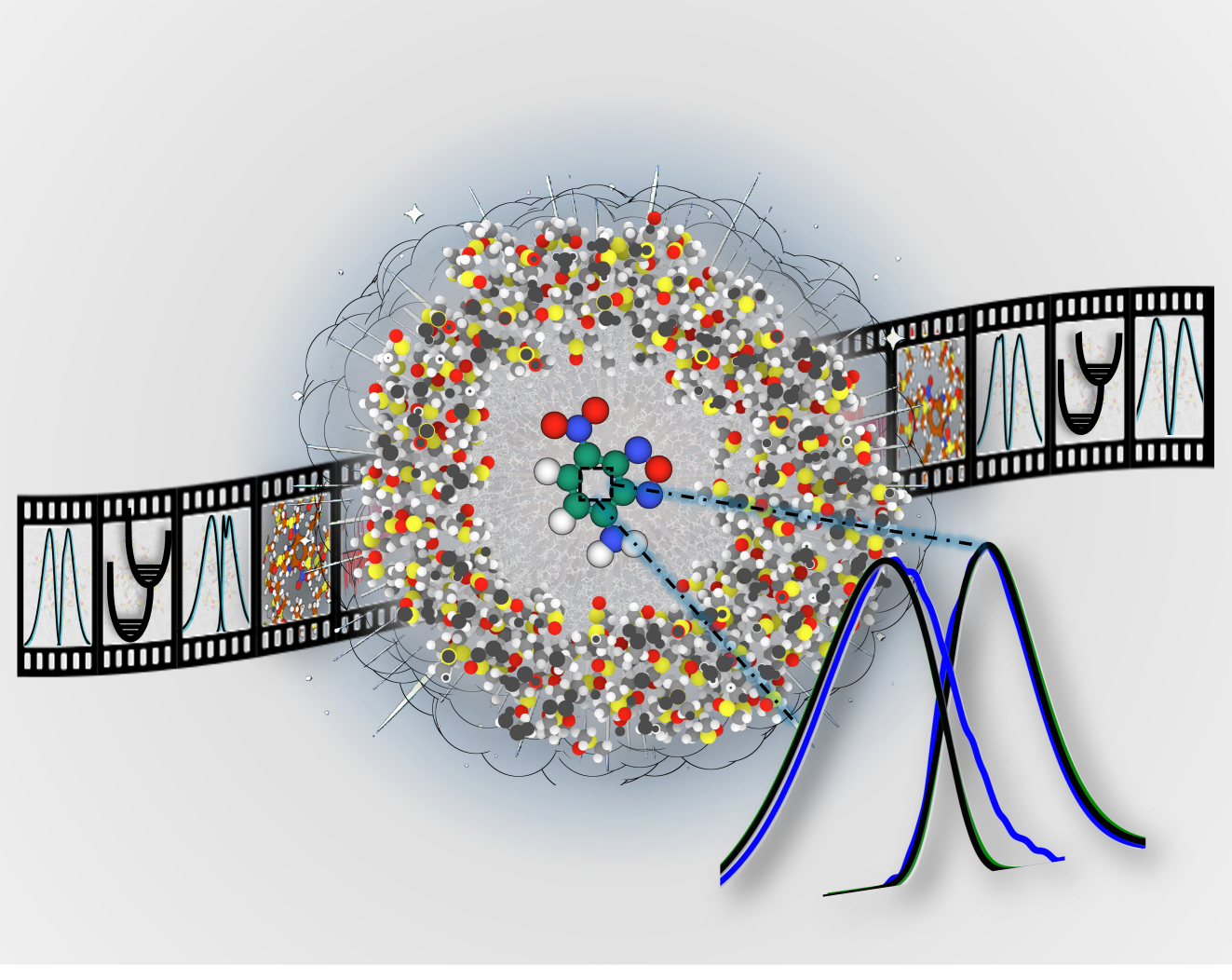

Computing Absorption and Fluorescence Spectra
Calculating absorption and fluroscence spectra of molecules in explicit environment using ensemble Franck-Condon methods
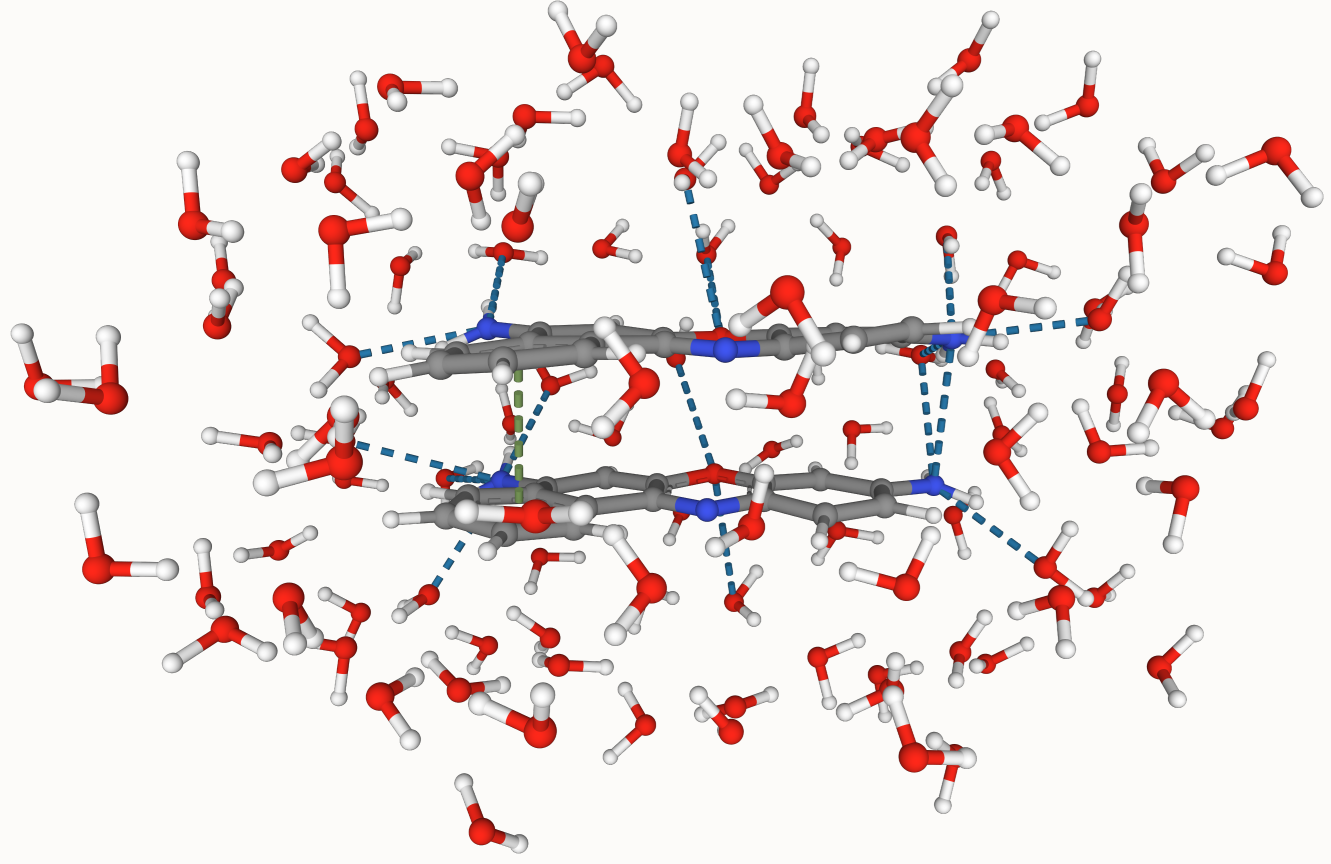
Automate QM/MM Sims
A Python program designed automate MD trajectories containing dyes in solvents to hybrid QM/MM simulations.
MolVizMan
An interactive Python GUI application allows you to visualize and manipulate molecules from an XYZ file.
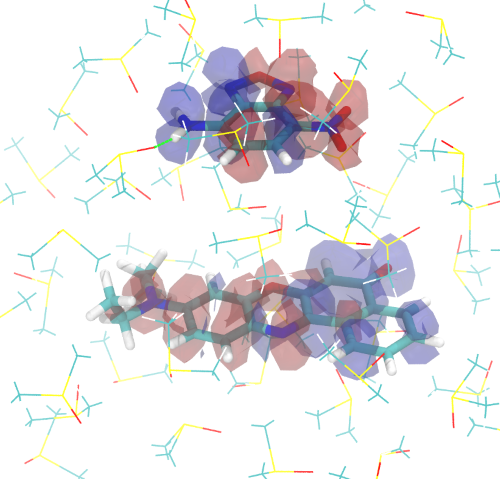
Excitonic Coupling
Computing Excitonic Coupling Between Molecules Using Various Excitonic Coupling Schemes

DOI2BibTex
Convert Digital Object Identifiers (DOI) into BibTeX entries with a simple, user-friendly web app built using Streamlit!
Technical Expertise
Combining quantum mechanics, molecular simulations, and machine learning for real-world impact
Drug Discovery
Virtual screening, docking, and ADMET prediction for therapeutic design
QM/MM & Spectroscopy
Hybrid quantum/classical methods for excited states and optical properties
Energy Materials
Periodic DFT for batteries, catalysts, and photovoltaic materials
Machine Learning
ML/DL models for property prediction, QSAR, and molecular generation
Programming & HPC
High-performance computing and GPU acceleration for large-scale simulations
Cheminformatics
Large-scale compound screening, similarity search, and QSAR modeling
Professional Experience
Research and development at the forefront of computational chemistry
Postdoctoral Research Associate
Los Alamos National Laboratory (LANL)
Advisor: Dr. Sergei Tretiak
Key Achievements:
- Developing deep learning-based surrogate models for non-adiabatic molecular dynamics simulations
- Established quantitative structure-property relationships (QSPR) in π-stacked molecular aggregates (perylene diimide, squaraine, and cyanine dyes)
- Generated foundational insights into the structural origins of chiroptical signals in chiral molecular aggregates
Research Focus:
Graduate Researcher
University of California, Merced
Computational Chemistry & Physics
Key Achievements:
- Extended ensemble Franck-Condon methods for accurate full stack UV-Vis spectroscopy predictions in solution (J. Chem. Phys. 2024)
- Co-authored Nature Communications publication on molecular polariton electroabsorption (16+ citations)
- Developed QM/MM automation tools reducing simulation setup time by 70%
Research Focus:
Computer-Aided Drug Design (CADD) Intern
Frontier Medicines
Computational Chemistry & Drug Discovery
Key Achievements:
- Developed Python pipelines to automate SMILES-to-desolvation energy calculations and conformational sampling for Bruton's Tyrosine Kinase (BTK) inhibitors, leveraging OpenMM and TeraChem (hybrid QM/MM interface) for significantly streamlined workflows
- Gained proficiency in unbiased and biased ligand-based docking (MOE), successfully deploying these techniques to perform protein-ligand docking for BTK inhibitors
- Built robust Protein-Ligand binding free energy calculation pipelines, utilizing hybrid QM/MM techniques, to enable accurate rank-ordering of Bruton's Tyrosine Kinase (BTK) inhibitors
Technologies:
Teaching & Mentorship
Graduate Mentoring
- • GradEXCEL Program mentor (2019-2022)
- • Supervised undergraduate researchers
- • Graduate Excel Peer Mentor Award
Teaching Development
- • Advanced Pedagogy certification
- • GROW TA Training Fellowship
- • Stanford Writing in Sciences course
🌟 Featured Publications
1. Covalent Control of Excitonic Interactions in Perylene Diimide Trimers: A Computational Study
Ajay Khanna, Jean-Huber Olivier, Sebastian Fernandez-Albertia, and Sergei Tretiak Nano Letters (2026) Top 10% Novelty
|| Comprehensive study investigating structural disorder effects on electronic and optical properties in π-stacked perylene diimide aggregates, providing insights for organic photovoltaic design.
2. Deconstructing Chirality: Probing Local and Nonlocal Effects in Azobenzene Derivatives with X-ray Circular Dichroism
Ajay Khanna, Victor M. Freixas, Lei Xu, Jérémy R. Rouxel, Niranjan Govind, Marco Garavelli, Shaul Mukamel, and Sergei Tretiak J. Phys. Chem. Lett. 16 (2025) Journal Cover Top 15% Novelty
|| Delivered foundational insights into the structural origins of chiroptical signals, leading to a mechanistic understanding crucial for the rational design of functional molecular machines. Accepted without revision and selected for journal cover.
3. Calculating Absorption and Fluorescence Spectra for Chromophores in Solution with Ensemble Franck-Condon Methods
Ajay Khanna, Sapana V. Shedge, Tim J. Zuehlsdorff, Christine M. Isborn J. Chem. Phys. (2024)Citations: 6 Top 5-25% Novelty
|| Novel computational approach for accurate prediction of absorption and fluorescence spectra in solution, advancing spectroscopic analysis techniques.
4. Molecular Polariton Electroabsorption
Chiao-Yu Cheng, Nina Krainova, Alyssa Brigeman, Ajay Khanna, Sapana Shedge, Christine Isborn, Joel Yuen-Zhou, Noel C. Giebink Nature Comm. (2022) Citations: 22
|| Novel study on molecular polariton electroabsorption, opening new avenues in optoelectronics and quantum technology.
5. Explicit Environmental and Vibronic Effects in Simulations of Linear and Nonlinear Optical Spectroscopy
Sapana V. Shedge, Tim J. Zuehlsdorff, Ajay Khanna, Stacey Conley, Christine M. Isborn J. Chem. Phys. (2021) Citations: 15
|| Comprehensive exploration of environmental and vibronic effects in optical spectroscopy simulations, enhancing accuracy in molecular property predictions.
6. Axial H-Bonding Solvent Controls Inhomogeneous Spectral Broadening, While Peripheral H-Bonding Solvent Controls Vibronic Broadening: Cresyl Violet in Methanol
Christopher A. Myers, Shao-Yu Lu, Sapana Shedge, Arthur Pyuskulyan, Katherine Donahoe, Ajay Khanna, Liang Shi, and Christine M. Isborn The Journal of Physical Chemistry B 2024 Citations: 6
|| Systematic dissection of solvent effects on spectral line shapes, improving simulation accuracy by identifying the critical role of specific molecular interactions.
7. Ligand Driven Electron Counting Rule Selection: A Case Study for Ge5R Complex
Rakesh Parida, G. Naaresh Reddy, Ajay Khanna, Gourisankar Roymahapatra, Santanab Giri Int. J. HIT. TRANSC: ECCN. Vol (2018)
|| Theoretical investigation of ligand-driven electron counting rules in germanium cluster complexes, contributing to understanding of main-group element chemistry.
Global Research Collaboration
Working with leading researchers and institutions worldwide to advance computational chemistry
My Research Journey
From first publication to cutting-edge postdoctoral research
First Publication
Ligand Driven Electron Counting Rule Selection (NIT Rourkela collaboration)
PublicationPh.D. Started
UC Merced - Computational Chemistry
🏆 Summer Research Fellowship
Summer Research Fellowship
UC Merced (2nd consecutive year)
AwardJ. Chem. Phys. Publication
Explicit Environmental Effects in Optical Spectroscopy
🎤 ACS Conference (Talk & Poster)
🎤 UC Merced Talk
Nature Communications
Molecular Polariton Electroabsorption (22 citations)
🏆 5 Awards & Fellowships:
• Graduate Excel Peer Mentor Award
• Graduate Fellowship Incentive Program
• GROW TA Training Fellowship
• Chemistry Travel Award
🎤 ACS Spring Conference (Talk)
CADD Intern - Frontier Medicines
Virtual screening, QM/MM, and drug design
🏆 Outstanding Graduate Student Award (UC
Merced)
💰 XSEDE/ACCESS HPC Grant ($5,263)
8000 GPU-hours + 3000 CPU-hours
Ph.D. Completed
UC Merced - Computational Chemistry
📄 2 Publications:
• J. Chem. Phys. (Franck-Condon Methods - 6 citations)
• J. Phys. Chem. B (Spectral Broadening - 6 citations)
🎤 WCTC Conference (Poster)
Postdoc - Los Alamos National Laboratory
Deep Learning for Non-Adiabatic MD & Chiroptical Spectroscopy
📄 2 Publications (2025):
• J. Phys. Chem. Lett. (Journal Cover + Top 15% Novelty)
• J. Phys. Chem. C. (Under Review)
🎤 ESP 2025 Conference (Poster)
Professional Certifications
Continuous learning and specialization in cutting-edge computational methods
Introduction to Cheminformatics and Medicinal Chemistry
Udemy
2024
Comprehensive training in computational methods for molecular representation, drug-target interactions, and structure-based drug design
Data Science with Python
Simplilearn
2022
Expertise in data analysis, visualization, and machine learning using Python ecosystem (NumPy, Pandas, Scikit-learn)
Fundamental of Accelerated Computing with CUDA Python
NVIDIA
2023
GPU-accelerated computing and parallel programming for high-performance scientific simulations
Tutorials

Running Classical MD Sims
Getting started with running classical molecular dynamics simulations using Amber24

Running AIMD Simulations
How to run ab initio molecular dynamics (AIMD) simulations in gas, implicit and explicit Enviroments

Everything TeraChem
TeraChem for electronic structure calculations: Gas, implicit and explicit environments

Everything Gaussian
Gaussian16 for electronic structure calculations: Gas, implicit and explicit environments

More Tutorials
Click For More Tutorials on different types of MD and AIMD sims. Electronic structure, data analysis, quality images etc